Drug design: Immunosuppressant developed on a computer
ETH Zurich scientists, using computer software they developed and in collaboration with German immunologists, have discovered a pharmaceutically active molecule that can halt the excessive production of an immune system messenger substance. A derivative of this molecule could one day help patients with an autoimmune disease.
Interferons are important messenger substances in human and animal immune systems. For example, one class of interferons, known as Type I interferons, are produced by cells of the immune system as a response to infections by pathogenic viruses or bacteria.
However, there are also illnesses in which interferon-producing cells are chronically activated. This can lead to a feedback effect and thus to overproduction of the messenger substances. As a consequence, the immune system runs out of control, causing a chronic inflammatory reaction. This happens for example in the case of systemic lupus erythematosus, a serious autoimmune disease.
Researchers led by Gisbert Schneider, Professor of Computer-Assisted Drug Design at the Institute of Pharmaceutical Sciences of ETH Zurich, together with Zoe Waibler’s work group at the Paul Ehrlich Institute in Langen (Germany), have discovered a pharmaceutically active candidate substance that specifically inhibits the secretion and activity of Type I interferons. In future this could help patients suffering from diseases such as systemic lupus erythematosus.
A prediction about the binding site
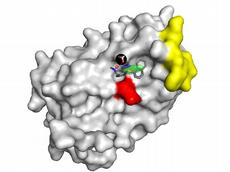
Gisbert Schneider and his colleagues at ETH Zurich did this by using a computer model to test the potential ability of a virtual collection of more than 500,000 substances to bind to Type I interferons. For this, the model used the three-dimensional structural information about the messenger substance, its receptor on the surface of immune cells, and the pharmaceutically active candidate substances from the virtual library. As Tim Geppert, ETH doctoral student and first author of the study explains, new software developed at ETH Zurich was used for this purpose. For two given molecules – in this case the messenger substance and its receptor – the software calculates where and how they are most likely to bind to one another and to interact. The software also predicted a binding site for druglike compounds, which might block the interaction between the messenger and its receptor.
Six molecules from the virtual library proved especially promising in this computer simulation. The ability of these compounds to bind to interferon was then examined in biophysical studies in the laboratory. Finally the efficacy of the most suitable substances was checked and confirmed in cell cultures by scientists at the Paul Ehrlich Institute. The researchers tested how well the pharmaceutically active candidate substances suppressed the feedback effect in interferon-producing cells.
More research is needed
However, the most effective molecule will not be directly usable as a new drug, because the cell culture experiments also proved it to be toxic to the cells at high doses. Therefore, in the next step, the scientists want to develop slightly modified derivatives of the newly discovered active substance that still have the desired activity but are non-toxic. Schneider says, “As this study has shown, early-phase drug discovery can profit enormously from close collaboration between scientists from different disciplines.” Chemical and bio-informatics played a decisive role in this endeavour.
Literature reference:
Geppert T, Bauer S, Hiss JA, Conrad E, Reutlinger M, Schneider P, Weisel M, Pfeiffer B, Altmann KH, Waibler Z, Schneider G: Immunosuppressive Small Molecule Discovered by Structure-Based Virtual Screening for Inhibitors of Protein-Protein Interactions. Angewandte Chemie International Edition 2011. doi: 10.1002/anie.201105901








READER COMMENTS